Neurons#
Neurons are the fundamental computational units in brainpy.state. This document explains how neurons work, what models are available, and how to use and create them.
Overview#
In brainpy.state, neurons model the dynamics of neural populations. Each neuron model:
Maintains membrane potential (voltage)
Integrates input currents
Generates spikes when threshold is crossed
Resets after spiking (various strategies)
All neuron models inherit from the base Neuron class and follow consistent interfaces.
Basic Usage#
Creating Neurons#
import numpy as np
import brainpy.state
import brainstate
import braintools
import saiunit as u
import jax.numpy as jnp
An NVIDIA GPU may be present on this machine, but a CUDA-enabled jaxlib is not installed. Falling back to cpu.
brainstate.environ.set(dt=0.1 * u.ms)
# Create a population of 100 LIF neurons
neurons = brainpy.state.LIF(
in_size=100,
V_rest=-65. * u.mV,
V_th=-50. * u.mV,
V_reset=-65. * u.mV,
tau=10. * u.ms
)
Initializing States#
Before simulation, initialize neuron states:
# Initialize all states to default values
brainstate.nn.init_all_states(neurons)
# Or with specific batch in_size
brainstate.nn.init_all_states(neurons, batch_size=32)
LIF(
in_size=(100,),
out_size=(100,),
spk_reset=soft,
spk_fun=ReluGrad(alpha=0.3, width=1.0),
R=Quantity(1., "ohm"),
tau=Quantity(10., "ms"),
V_th=Quantity(-50., "mV"),
V_rest=Quantity(-65., "mV"),
V_reset=Quantity(-65., "mV"),
V_initializer=Constant(value=0. mV),
V=HiddenState(
value=Quantity(~float32[32,100], "mV")
)
)
Running Neurons#
Update neurons by calling them with input current:
# Single time step - provide input for all neurons
# Create input current array matching neuron population in_size
input_current = jnp.ones(100) * 2.0 * u.nA # 100 neurons, each gets 2.0 nA
# Access results
voltage = neurons.V.value # Membrane potential
spikes = neurons.get_spike() # Spike output
print(f"Voltage shape: {voltage.shape}")
print(f"Spikes shape: {spikes.shape}")
Voltage shape: (32, 100)
Spikes shape: (32, 100)
Available Neuron Models#
For more neuron models, see the API Reference.
IF (Integrate-and-Fire)#
The simplest spiking neuron model.
Mathematical Model:
Spike condition: If \(V \geq V_{th}\), emit spike and reset.
Example:
# IF neuron - simple parameters
neuron = brainpy.state.IF(
in_size=100,
V_th=1. * u.mV, # Spike threshold
tau=20. * u.ms, # Membrane time constant
R=1. * u.ohm # Input resistance
)
# Initialize the neuron
import brainstate
brainstate.nn.init_all_states(neuron)
IF(
in_size=(100,),
out_size=(100,),
spk_reset=soft,
spk_fun=ReluGrad(alpha=0.3, width=1.0),
R=Quantity(1., "ohm"),
tau=Quantity(20., "ms"),
V_th=Quantity(1., "mV"),
V_initializer=Constant(value=0. mV),
V=HiddenState(
value=Quantity(~float32[100], "mV")
)
)
Parameters:
in_size: Number of neuronsV_rest: Resting potentialV_th: Spike thresholdV_reset: Reset potential after spiketau: Membrane time constantR: Input resistance
Use cases:
Simple rate coding
Fast simulations
Theoretical studies
LIF (Leaky Integrate-and-Fire)#
The most commonly used spiking neuron model.
Mathematical Model:
Spike condition: If \(V \geq V_{th}\), emit spike and reset.
Example:
neuron = brainpy.state.LIF(
in_size=100,
V_rest=-65. * u.mV,
V_th=-50. * u.mV,
V_reset=-65. * u.mV,
tau=10. * u.ms,
R=1. * u.ohm,
V_initializer=braintools.init.Normal(-65., 5., unit=u.mV)
)
# Initialize the neuron
brainstate.nn.init_all_states(neuron)
LIF(
in_size=(100,),
out_size=(100,),
spk_reset=soft,
spk_fun=ReluGrad(alpha=0.3, width=1.0),
R=Quantity(1., "ohm"),
tau=Quantity(10., "ms"),
V_th=Quantity(-50., "mV"),
V_rest=Quantity(-65., "mV"),
V_reset=Quantity(-65., "mV"),
V_initializer=Normal(mean=-65.0, std=5.0),
V=HiddenState(
value=Quantity(float32[100], "mV")
)
)
Parameters:
All IF parameters, plus:
V_initializer: How to initialize membrane potential
Key Features:
Leak toward resting potential
Realistic temporal integration
Well-studied dynamics
Use cases:
General spiking neural networks
Cortical neuron modeling
Learning and training
LIFRef (LIF with Refractory Period)#
LIF neuron with absolute refractory period.
Mathematical Model:
Same as LIF, but after spiking:
Neuron is “frozen” for refractory period
No integration during refractory period
More biologically realistic
Example:
neuron = brainpy.state.LIFRef(
in_size=100,
V_rest=-65. * u.mV,
V_th=-50. * u.mV,
V_reset=-65. * u.mV,
tau=10. * u.ms,
tau_ref=2. * u.ms, # Refractory period
R=1. * u.ohm
)
# Initialize the neuron
brainstate.nn.init_all_states(neuron)
LIFRef(
in_size=(100,),
out_size=(100,),
spk_reset=soft,
spk_fun=ReluGrad(alpha=0.3, width=1.0),
R=Quantity(1., "ohm"),
tau=Quantity(10., "ms"),
tau_ref=Quantity(2., "ms"),
V_th=Quantity(-50., "mV"),
V_rest=Quantity(-65., "mV"),
V_reset=Quantity(-65., "mV"),
V_initializer=Constant(value=0. mV),
V=HiddenState(
value=Quantity(~float32[100], "mV")
),
last_spike_time=ShortTermState(
value=Quantity(~float32[100], "ms")
)
)
Additional Parameters:
tau_ref: Refractory period duration
Key Features:
Absolute refractory period
Prevents immediate re-firing
More realistic spike timing
Use cases:
Precise temporal coding
Biological realism
Rate regulation
ALIF (Adaptive Leaky Integrate-and-Fire)#
LIF with spike-frequency adaptation.
Mathematical Model:
When spike occurs: \(w \leftarrow w + \beta\)
Example:
neuron = brainpy.state.ALIF(
in_size=100,
V_rest=-65. * u.mV,
V_th=-50. * u.mV,
V_reset=-65. * u.mV,
tau=10. * u.ms,
tau_a=200. * u.ms, # Adaptation time constant
beta=0.1 * u.nA, # Spike-triggered adaptation
R=1. * u.ohm
)
# Initialize the neuron
brainstate.nn.init_all_states(neuron)
ALIF(
in_size=(100,),
out_size=(100,),
spk_reset=soft,
spk_fun=ReluGrad(alpha=0.3, width=1.0),
R=Quantity(1., "ohm"),
tau=Quantity(10., "ms"),
tau_a=Quantity(200., "ms"),
V_th=Quantity(-50., "mV"),
V_reset=Quantity(-65., "mV"),
V_rest=Quantity(-65., "mV"),
beta=Quantity(0.1, "nA"),
V_initializer=Constant(value=0. mV),
a_initializer=Constant(value=0.0),
V=HiddenState(
value=Quantity(~float32[100], "mV")
),
a=HiddenState(
value=ShapedArray(float32[100], weak_type=True)
)
)
Additional Parameters:
tau_w: Adaptation time constantbeta: Adaptation increment per spike
Key Features:
Spike-frequency adaptation
Reduced firing with sustained input
More complex dynamics
Use cases:
Cortical neuron modeling
Sensory adaptation
Complex temporal patterns
Reset Modes#
BrainPy supports different reset behaviors after spiking:
Soft Reset (Default)#
Subtract threshold from membrane potential:
neuron = brainpy.state.LIF(
in_size=100,
V_rest=-65. * u.mV,
V_th=-50. * u.mV,
V_reset=-65. * u.mV,
tau=10. * u.ms,
spk_reset='soft'
)
brainstate.nn.init_all_states(neuron)
LIF(
in_size=(100,),
out_size=(100,),
spk_reset=soft,
spk_fun=ReluGrad(alpha=0.3, width=1.0),
R=Quantity(1., "ohm"),
tau=Quantity(10., "ms"),
V_th=Quantity(-50., "mV"),
V_rest=Quantity(-65., "mV"),
V_reset=Quantity(-65., "mV"),
V_initializer=Constant(value=0. mV),
V=HiddenState(
value=Quantity(~float32[100], "mV")
)
)
Properties:
Preserves extra charge above threshold
Allows rapid re-firing
Common in machine learning
Hard Reset#
Reset to fixed potential:
neuron = brainpy.state.LIF(
in_size=100,
V_rest=-65. * u.mV,
V_th=-50. * u.mV,
V_reset=-65. * u.mV,
tau=10. * u.ms,
spk_reset='hard'
)
brainstate.nn.init_all_states(neuron)
LIF(
in_size=(100,),
out_size=(100,),
spk_reset=hard,
spk_fun=ReluGrad(alpha=0.3, width=1.0),
R=Quantity(1., "ohm"),
tau=Quantity(10., "ms"),
V_th=Quantity(-50., "mV"),
V_rest=Quantity(-65., "mV"),
V_reset=Quantity(-65., "mV"),
V_initializer=Constant(value=0. mV),
V=HiddenState(
value=Quantity(~float32[100], "mV")
)
)
Properties:
Discards extra charge
More biologically realistic
Prevents immediate re-firing
Choosing Reset Mode#
Soft reset: Machine learning, rate coding, fast oscillations
Hard reset: Biological modeling, temporal coding, realism
Spike Functions#
For training spiking neural networks, use surrogate gradients:
neuron = brainpy.state.LIF(
in_size=100,
V_rest=-65. * u.mV,
V_th=-50. * u.mV,
V_reset=-65. * u.mV,
tau=10. * u.ms,
spk_fun=braintools.surrogate.ReluGrad()
)
brainstate.nn.init_all_states(neuron)
LIF(
in_size=(100,),
out_size=(100,),
spk_reset=soft,
spk_fun=ReluGrad(alpha=0.3, width=1.0),
R=Quantity(1., "ohm"),
tau=Quantity(10., "ms"),
V_th=Quantity(-50., "mV"),
V_rest=Quantity(-65., "mV"),
V_reset=Quantity(-65., "mV"),
V_initializer=Constant(value=0. mV),
V=HiddenState(
value=Quantity(~float32[100], "mV")
)
)
Available surrogate functions:
ReluGrad(): ReLU-like gradientSigmoidGrad(): Sigmoid-like gradientGaussianGrad(): Gaussian-like gradientSuperSpike(): SuperSpike surrogate
See Tutorial 3 for training details.
Advanced Features#
Initialization Strategies#
Different ways to initialize membrane potential:
# Constant initialization
neuron1 = brainpy.state.LIF(
in_size=100,
V_rest=-65. * u.mV,
V_th=-50. * u.mV,
tau=10. * u.ms,
V_initializer=braintools.init.Constant(-65., unit=u.mV)
)
brainstate.nn.init_all_states(neuron1)
# Normal distribution
neuron2 = brainpy.state.LIF(
in_size=100,
V_rest=-65. * u.mV,
V_th=-50. * u.mV,
tau=10. * u.ms,
V_initializer=braintools.init.Normal(-65., 5., unit=u.mV)
)
brainstate.nn.init_all_states(neuron2)
# Uniform distribution
neuron3 = brainpy.state.LIF(
in_size=100,
V_rest=-65. * u.mV,
V_th=-50. * u.mV,
tau=10. * u.ms,
V_initializer=braintools.init.Uniform(-70., -60., unit=u.mV)
)
brainstate.nn.init_all_states(neuron3)
LIF(
in_size=(100,),
out_size=(100,),
spk_reset=soft,
spk_fun=ReluGrad(alpha=0.3, width=1.0),
R=Quantity(1., "ohm"),
tau=Quantity(10., "ms"),
V_th=Quantity(-50., "mV"),
V_rest=Quantity(-65., "mV"),
V_reset=Quantity(0., "mV"),
V_initializer=Uniform(low=-70.0, high=-60.0),
V=HiddenState(
value=Quantity(float32[100], "mV")
)
)
Accessing Neuron States#
# Membrane potential (with units)
voltage = neuron.V.value # Quantity with units
# Spike output (binary or real-valued)
spikes = neuron.get_spike()
# Access underlying array (without units)
voltage_array = neuron.V.value.to_decimal(u.mV)
Batched Simulation#
Simulate multiple trials in parallel:
# Initialize with batch dimension
brainstate.nn.init_all_states(neuron, batch_size=32)
# Input shape: (batch_in_size, in_size)
# For 32 batches of 100 neurons each
input_current = jnp.ones((32, 100)) * 2.0 * u.nA
neuron(input_current)
# Output shape: (batch_in_size, in_size)
spikes = neuron.get_spike()
print(f"Spikes shape: {spikes.shape}") # Should be (32, 100)
Spikes shape: (32, 100)
Complete Example#
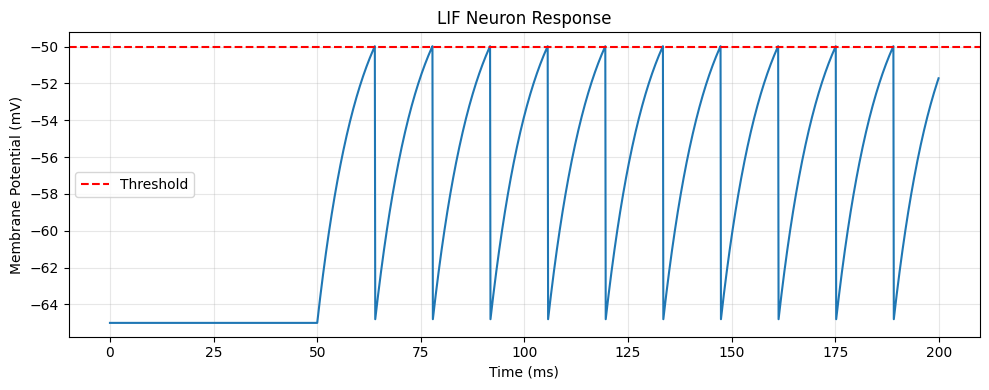
Here’s a complete example simulating a LIF neuron:
import matplotlib.pyplot as plt
# Set time step
brainstate.environ.set(dt=0.1 * u.ms)
# Create neuron
neuron = brainpy.state.LIF(
in_size=1,
V_rest=-65. * u.mV,
V_th=-50. * u.mV,
V_reset=-65. * u.mV,
tau=10. * u.ms,
spk_reset='hard'
)
# Initialize
brainstate.nn.init_all_states(neuron)
# Simulation parameters
duration = 200. * u.ms
dt = brainstate.environ.get_dt()
times = u.math.arange(0. * u.ms, duration, dt)
# Input current (step input)
def get_input():
t = brainstate.environ.get('t')
return u.math.where(
t > 50*u.ms,
jnp.ones(1) * 20.0 * u.mA, # Array of in_size 1
jnp.zeros(1) * u.mA, # Array of in_size 1
)
def step_run(i, t):
with brainstate.environ.context(i=i, t=t):
neuron(get_input())
return neuron.V.value, neuron.get_spike()
# Run simulation
voltages, spikes = brainstate.transform.for_loop(step_run, jnp.arange(times.size), times)
# Plot results
voltages = u.math.asarray(voltages)
times_plot = times.to_decimal(u.ms)
voltages_plot = voltages.to_decimal(u.mV).squeeze() # Remove in_size dimension
plt.figure(figsize=(10, 4))
plt.plot(times_plot, voltages_plot)
plt.axhline(y=-50, color='r', linestyle='--', label='Threshold')
plt.xlabel('Time (ms)')
plt.ylabel('Membrane Potential (mV)')
plt.title('LIF Neuron Response')
plt.legend()
plt.grid(True, alpha=0.3)
plt.tight_layout()
plt.show()
Creating Custom Neurons#
You can create custom neuron models by inheriting from Neuron:
from brainpy.state import Neuron
class MyNeuron(Neuron):
def __init__(self, in_size, tau, V_th, **kwargs):
super().__init__(in_size, **kwargs)
# Store parameters
self.tau = tau
self.V_th = V_th
# Initialize states
self.V = brainstate.ShortTermState(
braintools.init.Constant(0., unit=u.mV)(in_size)
)
self.spike = brainstate.ShortTermState(
jnp.zeros(in_size)
)
def update(self, x):
# Get time step
dt = brainstate.environ.get_dt()
# Update membrane potential (custom dynamics)
dV = (-self.V.value + x) / self.tau * dt
V_new = self.V.value + dV
# Check for spikes
spike = (V_new >= self.V_th).astype(float)
# Reset
V_new = jnp.where(spike > 0, 0. * u.mV, V_new)
# Update states
self.V.value = V_new
self.spike.value = spike
return spike
def get_spike(self):
return self.spike.value
Usage:
neuron = MyNeuron(in_size=100, tau=10*u.ms, V_th=1*u.mV)
brainstate.nn.init_all_states(neuron)
# Create appropriate input current
input_current = jnp.ones(100) * 0.5 * u.nA
Performance Tips#
Use JIT compilation for repeated simulations:
@brainstate.transform.jit
def simulate_step(input):
neuron(input)
return neuron.V.value
Batch multiple trials for parallelism:
brainstate.nn.init_all_states(neuron, batch_size=100)
MyNeuron(
in_size=(100,),
out_size=(100,),
spk_reset=soft,
spk_fun=InvSquareGrad(alpha=100.0),
tau=Quantity(10, "ms"),
V_th=Quantity(1, "mV"),
V=ShortTermState(
value=Quantity(~float32[100], "mV")
),
spike=ShortTermState(
value=ShapedArray(float32[100])
)
)
Use appropriate data types:
# Float32 is usually sufficient and faster
brainstate.environ.set(precision=32)
Use soft reset for higher firing rates:
# Use soft reset for higher firing rates
neuron = brainpy.state.LIF(100, tau=10*u.ms, spk_reset='soft')
Use hard reset for precise spike timing:
# Use refractory period for precise timing
neuron = brainpy.state.LIFRef(
in_size=100,
V_rest=-65. * u.mV,
V_th=-50. * u.mV,
V_reset=-65. * u.mV,
tau=10. * u.ms,
tau_ref=2. * u.ms,
spk_reset='hard'
)
Use refractory period for precise timing
neuron = brainpy.state.LIFRef(
100,
tau=10*u.ms,
tau_ref=2*u.ms,
spk_reset='hard'
)
Adaptation creates bursting patterns
neuron = brainpy.state.ALIF(
in_size=100,
V_rest=-65. * u.mV,
V_th=-50. * u.mV,
V_reset=-65. * u.mV,
tau=10. * u.ms,
tau_a=200. * u.ms,
spk_reset='soft'
)
brainstate.nn.init_all_states(neuron)
# Adaptation creates bursting patterns
neuron = brainpy.state.ALIF(
100,
tau=10*u.ms,
tau_a=200*u.ms,
beta=0.01,
spk_reset='soft'
)
Summary#
Neurons in brainpy.state:
✅ Multiple models: IF, LIF, LIFRef, ALIF
✅ Physical units: All parameters with proper units
✅ Flexible reset: Soft or hard reset modes
✅ Training-ready: Surrogate gradients for learning
✅ High performance: JIT compilation and batching
✅ Extensible: Easy to create custom models
Next Steps#
Learn about synapses to connect neurons
Explore projections for network connectivity
Follow tutorials for hands-on practice
See docs for network examples
