Simulation and Analysis of Excitatory-Inhibitory Neuronal Networks (E-I Networks)#
This example demonstrates the implementation of a classical excitatory-inhibitory (E-I) neuronal network using the braincell framework. By constructing an E-I network composed of Hodgkin-Huxley model neurons, you will learn how to implement hierarchical modeling from single neurons to networks in braincell, and analyze the spike dynamics of the network.
Preparation#
First, ensure that the necessary libraries (braincell, brainstate, brainunit, matplotlib) are installed, and import the required modules:
import brainpy
import brainunit as u
import matplotlib.pyplot as plt
import brainstate
import braincell
Code Explanation#
Parameter Definition#
First, define the key biophysical parameters, which determine the electrophysiological properties of the neurons and the network size:
# Neuron action potential firing threshold
V_th = -20. * u.mV
# Neuron membrane area
area = 20000 * u.um **2
area = area.in_unit(u.cm** 2)
# Membrane capacitance
Cm = (1 * u.uF * u.cm ** -2) * area # Total capacitance = specific capacitance × area
Define HH single neuron model#
Use SingleCompartment to construct a single neuron based on the HH model, including sodium channel INa, potassium channel IK, and leak current IL. These channels jointly determine the firing properties of the neuron:
class HH(braincell.SingleCompartment):
def __init__(self, in_size):
# Initialize single-compartment neuron
super().__init__(in_size, C=Cm, solver='ind_exp_euler')
# Sodium channel (INa)
self.na = braincell.ion.SodiumFixed(in_size, E=50. * u.mV)
self.na.add_elem(
# Maximum conductance
INa=braincell.channel.INa_TM1991(in_size, g_max=(100. * u.mS * u.cm **-2) * area, V_sh=-63. * u.mV)
)
# Potassium channel (IK)
self.k = braincell.ion.PotassiumFixed(in_size, E=-90 * u.mV)
self.k.add_elem(
# Maximum conductance
IK=braincell.channel.IK_TM1991(in_size, g_max=(30. * u.mS * u.cm** -2) * area, V_sh=-63. * u.mV)
)
# Leak current (IL)
self.IL = braincell.channel.IL(
in_size,
E=-60. * u.mV,
g_max=(5. * u.nS * u.cm **-2) * area # Maximum conductance
)
Define E-I Network: Connections Between Excitatory and Inhibitory Neurons#
Construct a network composed of excitatory (E) and inhibitory (I) neurons to simulate the commonly observed E-I balance mechanism in cortical networks:
class EINet(brainstate.nn.Module):
def __init__(self):
super().__init__()
# Network scale
self.n_exc = 3200
self.n_inh = 800
self.num = self.n_exc + self.n_inh # Total number of neurons: 4000
# Initialize neuron population
self.N = HH(self.num)
# Excitatory synaptic projection
self.E = brainpy.state.AlignPostProj(
# Connection rule
comm=brainstate.nn.EventFixedProb(
self.n_exc, self.num, conn_num=0.02, # Connection probability
conn_weight=6. * u.nS # Synaptic weight
),
# Synaptic dynamics
syn=brainpy.state.Expon(self.num, tau=5. * u.ms),
# Postsynaptic effect
out=brainpy.state.COBA(E=0. * u.mV),
post=self.N # Projection target is neuron population N
)
# Inhibitory synaptic projection
self.I = brainpy.state.AlignPostProj(
# Connection rule
comm=brainstate.nn.EventFixedProb(
self.n_inh, self.num, conn_num=0.02,
conn_weight=67. * u.nS
),
# Synaptic dynamics
syn=brainpy.state.Expon(self.num, tau=10. * u.ms),
# Postsynaptic effect
out=brainpy.state.COBA(E=-80. * u.mV),
post=self.N # Projection target is neuron population N
)
def update(self, t):
# Define network update rules over time
with brainstate.environ.context(t=t):
# Get spike signals at current time
spk = self.N.spike.value
# Spikes from excitatory neurons drive excitatory synaptic projection
self.E(spk[:self.n_exc])
# Spikes from inhibitory neurons drive inhibitory synaptic projection
self.I(spk[self.n_exc:])
# Neurons update their states after receiving synaptic inputs, return new spike signals
spk = self.N(0. * u.nA)
Run network simulation#
Initialize the network and run the simulation, recording the spike activity of neurons at each time point:
# Initialize E-I network
net = EINet()
brainstate.nn.init_all_states(net) # Initialize the states of all neurons and synapses in the network
# Set simulation parameters and run
with brainstate.environ.context(dt=0.1 * u.ms): # Time step
# Generate simulation time sequence
times = u.math.arange(0. * u.ms, 100. * u.ms, brainstate.environ.get_dt())
# Loop to update network states
spikes = brainstate.transform.for_loop(
net.update, times,
pbar=brainstate.transform.ProgressBar(10) # Display progress bar
)
D:\Document\PyCharm\Project\braincell(collaborator)\.venv\Lib\site-packages\braintools\surrogate.py:72: UserWarning: Explicitly requested dtype float64 requested in asarray is not available, and will be truncated to dtype float32. To enable more dtypes, set the jax_enable_x64 configuration option or the JAX_ENABLE_X64 shell environment variable. See https://github.com/jax-ml/jax#current-gotchas for more.
z = jnp.asarray(x >= 0, dtype=x.dtype)
Running for 1,000 iterations: 100%|██████████| 1000/1000 [00:00<00:00, 13624.06it/s]
Visualize network spike activity#
Plot the spike data as a raster plot to intuitively show the firing patterns of neurons over time:
# Extract spike times and neuron indices
t_indices, n_indices = u.math.where(spikes)
# Plot raster plot
plt.scatter(times[t_indices], n_indices, s=1)
plt.xlabel('Time (ms)') # X-axis: time
plt.ylabel('Neuron index') # Y-axis: neuron index
plt.show()
# In this example, visualization cannot be completed after interruption in Jupyter, so temporarily keep the simulation results!
---------------------------------------------------------------------------
TypeError Traceback (most recent call last)
Cell In[24], line 2
1 # Extract spike times and neuron indices
----> 2 t_indices, n_indices = u.math.where(spikes)
4 # Plot raster plot
5 plt.scatter(times[t_indices], n_indices, s=1)
File D:\Document\PyCharm\Project\braincell(collaborator)\.venv\Lib\site-packages\saiunit\math\_fun_keep_unit.py:3305, in where(condition, x, y, size, fill_value)
3303 if x is None and y is None:
3304 assert not isinstance(fill_value, Quantity), "fill_value should not be a Quantity."
-> 3305 return jnp.where(condition, size=size, fill_value=fill_value)
3307 assert size is None and fill_value is None, "size and fill_value are only supported when x and y are not None."
3308 if isinstance(x, Quantity) and isinstance(y, Quantity):
File D:\Document\PyCharm\Project\braincell(collaborator)\.venv\Lib\site-packages\jax\_src\numpy\lax_numpy.py:2771, in where(condition, x, y, size, fill_value)
2711 """Select elements from two arrays based on a condition.
2712
2713 JAX implementation of :func:`numpy.where`.
(...) 2768 Array([0, 0, 0, 0, 0, 5, 6, 7, 8, 9], dtype=int32)
2769 """
2770 if x is None and y is None:
-> 2771 util.check_arraylike("where", condition)
2772 return nonzero(condition, size=size, fill_value=fill_value)
2773 else:
File D:\Document\PyCharm\Project\braincell(collaborator)\.venv\Lib\site-packages\jax\_src\numpy\util.py:183, in check_arraylike(fun_name, emit_warning, stacklevel, *args)
180 warnings.warn(msg + " In a future JAX release this will be an error.",
181 category=DeprecationWarning, stacklevel=stacklevel)
182 else:
--> 183 raise TypeError(msg.format(fun_name, type(arg), pos))
TypeError: where requires ndarray or scalar arguments, got <class 'NoneType'> at position 0.
Results Interpretation#
After running the code, you will see a raster plot, as shown below:
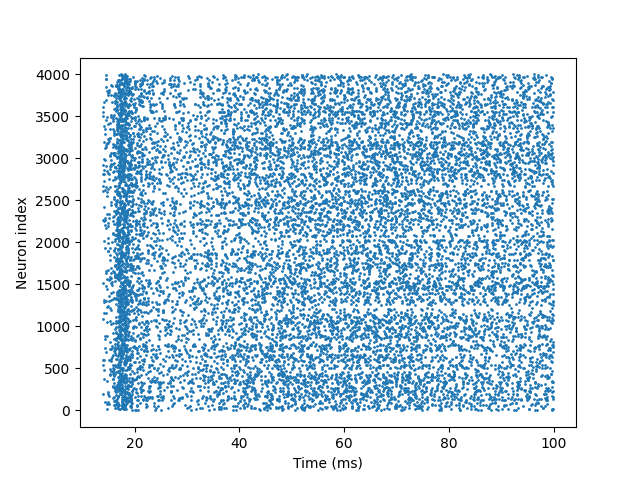
Each point represents a spike fired by a neuron at a specific time. Typical dynamics of an E-I network have the following characteristics:
Asynchronous firing: Neurons fire at dispersed times without obvious synchronous rhythms.
Sparse activity: Most neurons fire only a few times within 100 ms.
No burst synchronization: Rapid feedback from inhibitory synapses prevents large-scale synchronous bursting.
These features indicate that the E-I network maintains a stable and physiologically realistic activity pattern through dynamic balance between excitatory and inhibitory neurons.
Extended Exercises#
Adjust inhibitory synaptic weights and observe whether the network exhibits excessive synchronization or bursting activity.
Increase the proportion of excitatory neurons to analyze the impact of E-I imbalance on network dynamics.
Extend the simulation duration to see if the network maintains stable asynchronous activity.
Through these extensions, you can gain deeper insights into the critical role of E-I balance in maintaining neural network function, and the flexibility of braincell in modeling complex networks.
