Overview#
Welcome to the world of braincell!
This section provides a brief introduction to some of the key concepts and modeling features of the braincell framework.
braincell is a high-performance computing framework specifically designed for neuron modeling, built on top of JAX and brainstate. It offers a comprehensive toolchain for neuroscientists, computational neuroscientists, and neuromorphic computing engineers to construct, simulate, and optimize electrophysiologically precise models ranging from ion channels to multi-scale neural networks. The framework integrates advanced features such as modern hardware acceleration and automatic differentiation. The following tutorials will walk you through its layered architecture and core functionalities, helping you quickly understand the design logic behind braincell.
import brainunit as u
import brainpy
import braincell
import brainstate
import matplotlib.pyplot as plt
Hierarchical Architecture#
In neuron modeling, models are typically divided into the following major levels based on structural complexity and functional hierarchy:
ChannelIonCell
braincell focuses on precise neuronal dynamics modeling at these three levels.
Channel: Ion channels on the membrane, controlling the permeability of specific ions and determining the flow of ionic currents.Ion: The fundamental charged particles, such as Na⁺, K⁺, and Ca²⁺, that drive changes in membrane potential.Cell: The basic modeling unit of a single neuron, integrating multiple channels to produce membrane potential dynamics.
Taking the Hodgkin–Huxley (HH) neuron as an example, the relationship among these three levels can be illustrated as follows:
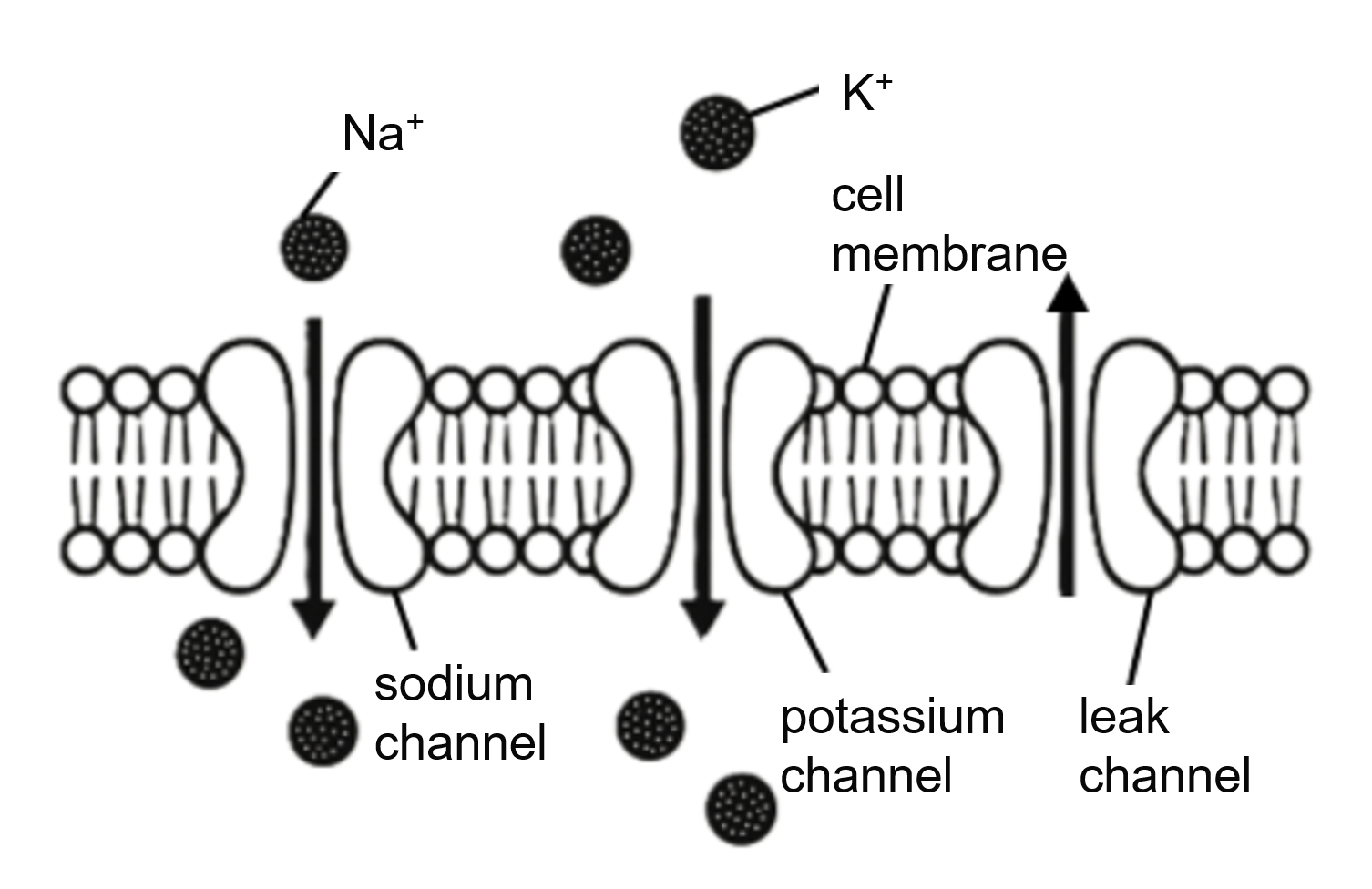
From this diagram, it is clear that Channel, Ion, and Cell are tightly interconnected. Biologically, these three levels form the fundamental mechanisms underlying neuronal electrical activity:
A neuron
Cellis surrounded by a cell membrane populated with various ionChannels.Ion
Channels regulate the flow of differentIons across the membrane.The movement of
Ions changes the potential difference across the membrane, thereby driving neuronal electrical activity.
Together, these levels provide a complete modeling pathway from microscopic mechanisms to macroscopic dynamics of neurons.
Core Features#
The main features of braincell include:
Ion channel modeling: Support for constructing electrophysiologically precise ion channel models using modular components.
Ion modeling: Support for modular construction of ion dynamics models.
Neuron modeling: Support for single-compartment and multi-compartment neuron models based on the HH framework.
Differential equation solvers: High-performance solvers built on JAX.
In the following sections, we will explore how to use and optimize these features, helping you build models ranging from complete neurons down to fine-grained ion mechanisms, and fully leverage the modeling capabilities of braincell.
Single-Compartment Neuron Modeling#
We now turn to how braincell can be used to construct neuron models.
Below is an example of building an HH neuron using braincell:
class HH(braincell.SingleCompartment):
def __init__(self, in_size):
super().__init__(in_size, C=Cm, solver='ind_exp_euler')
self.na = braincell.ion.SodiumFixed(in_size, E=50. * u.mV)
self.na.add(
INa=braincell.channel.INa_TM1991(in_size, g_max=(100. * u.mS * u.cm **-2) * area, V_sh=-63. * u.mV)
)
self.k = braincell.ion.PotassiumFixed(in_size, E=-90 * u.mV)
self.k.add(
IK=braincell.channel.IK_TM1991(in_size, g_max=(30. * u.mS * u.cm** -2) * area, V_sh=-63. * u.mV)
)
self.IL = braincell.channel.IL(
in_size,
E=-60. * u.mV,
g_max=(5. * u.nS * u.cm **-2) * area
)
We construct the HH neuron model by inheriting from SingleCompartment, which represents a single-compartment neuron.
SingleCompartment itself is the base class for all single-compartment neurons and is derived from HHTypedNeuron.
Through HHTypedNeuron, both single-compartment neurons (SingleCompartment) and multi-compartment neurons (MultiCompartment) can be modeled.
SingleCompartment provides built-in interfaces for membrane potential while supporting the integration of multiple Channels.
For example, INa_TM1991, IK_TM1991, and IL are concrete subclasses of the Channel class.
Taking INa_TM1991 as an example, it inherits from the sodium channel base class SodiumChannel, and thus provides the relevant interfaces.
From this example, we can see that modeling a neuron with braincell requires only the composition of the relevant classes at the three levels: Cell, Channel, and Ion.
The inheritance relationships within these three levels are deliberately kept simple in braincell.
The neuron Cell regulates the flow of different Ions, and each Ion is controlled by its corresponding Channel.
In the HH neuron model above:
The neuron includes sodium ions (implemented as
SodiumFixed) and potassium ions (implemented asPotassiumFixed).Sodium ions determine the behavior of the sodium channel
INa_TM1991.Potassium ions determine the behavior of the potassium channel
IK_TM1991.In addition,
ILrepresents a leakage channel, which is not associated with any ion.
This design makes neuron model construction both flexible and modular, facilitating extension and modification.
Neural Network Modeling#
Building upon single-neuron modeling in braincell, we can further construct neural network models, seamlessly integrating with the network modeling capabilities of BrainState.
Returning to our example, we can use the constructed HH neuron model to build networks for practical applications, such as an excitatory–inhibitory (E–I) network:
V_th = -20. * u.mV
area = 20000 * u.um ** 2
area = area.in_unit(u.cm ** 2)
Cm = (1 * u.uF * u.cm ** -2) * area # Membrane Capacitance [pF]
class EINet(brainstate.nn.Module):
def __init__(self):
super().__init__()
self.n_exc = 3200
self.n_inh = 800
self.num = self.n_exc + self.n_inh
self.N = HH(self.num)
self.E = brainpy.state.AlignPostProj(
comm=brainstate.nn.EventFixedProb(self.n_exc, self.num, conn_num=0.02, conn_weight=6. * u.nS),
syn=brainpy.state.Expon(self.num, tau=5. * u.ms),
out=brainpy.state.COBA(E=0. * u.mV),
post=self.N
)
self.I = brainpy.state.AlignPostProj(
comm=brainstate.nn.EventFixedProb(self.n_inh, self.num, conn_num=0.02, conn_weight=67. * u.nS),
syn=brainpy.state.Expon(self.num, tau=10. * u.ms),
out=brainpy.state.COBA(E=-80. * u.mV),
post=self.N
)
def update(self, t):
with brainstate.environ.context(t=t):
spk = self.N.spike.value
self.E(spk[:self.n_exc])
self.I(spk[self.n_exc:])
spk = self.N(0. * u.nA)
return spk
# network
net = EINet()
_ = brainstate.nn.init_all_states(net)
D:\Document\PyCharm\Project\braincell(collaborator)\.venv\Lib\site-packages\braintools\surrogate.py:72: UserWarning: Explicitly requested dtype float64 requested in asarray is not available, and will be truncated to dtype float32. To enable more dtypes, set the jax_enable_x64 configuration option or the JAX_ENABLE_X64 shell environment variable. See https://github.com/jax-ml/jax#current-gotchas for more.
z = jnp.asarray(x >= 0, dtype=x.dtype)
# simulation
with brainstate.environ.context(dt=0.1 * u.ms):
times = u.math.arange(0. * u.ms, 100. * u.ms, brainstate.environ.get_dt())
spikes = brainstate.transform.for_loop(net.update, times, pbar=brainstate.transform.ProgressBar(10))
Running for 1,000 iterations: 100%|██████████| 1000/1000 [00:00<00:00, 9927.91it/s]
# visualization
t_indices, n_indices = u.math.where(spikes)
plt.scatter(times[t_indices], n_indices, s=1)
plt.xlabel('Time (ms)')
plt.ylabel('Neuron index')
plt.show()
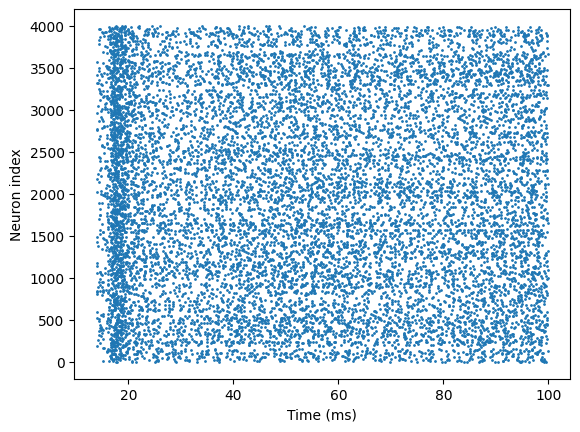
In this way, we have used braincell to model the HH neuron and implemented a complete E–I network!
